T4cluster is an R package designed as a computational toolkit with comprehensive coverage in relevant topics around the study of cluster analysis. It contains several classes of algorithms for
and other utility functions for further use. If you request additional functionalities or have suggestions, please contact maintainer.
You can install the released version of T4cluster from CRAN with:
install.packages("T4cluster")And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("kisungyou/T4cluster")T4cluster offers a variety of clustering algorithms
in common interface. In this example, we show a basic pipeline with
smiley dataset, which can be generated as follows;
# load the library
library(T4cluster)
# generate the data
smiley = T4cluster::genSMILEY(n=200)
data = smiley$data
label = smiley$label
# visualize
plot(data, pch=19, col=label, xlab="", ylab="", main="SMILEY Data")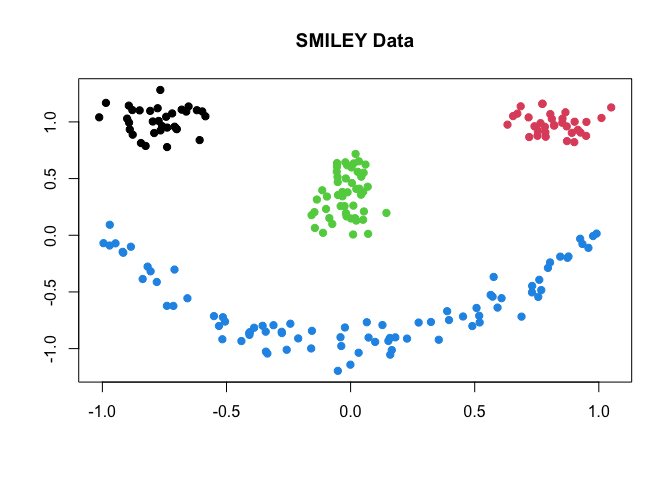
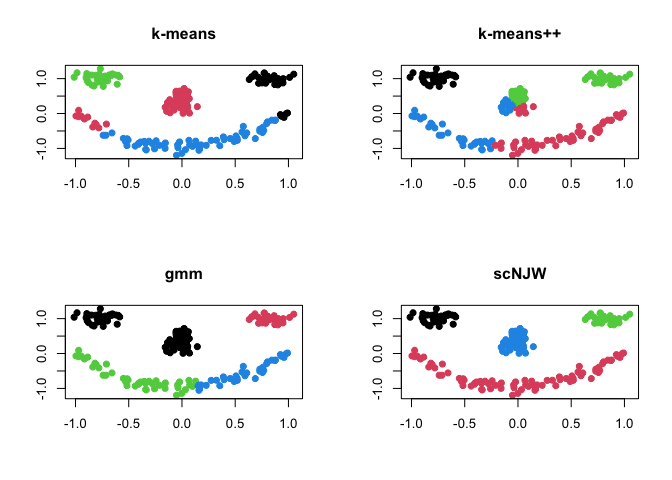
where each component of the face is considered as one cluster - the
data has 4 clusters. Here, we compare 4 different methods; (1)
k-means (kmeans), (2) k-means++
(kmeanspp), (3) gaussian mixture model (gmm),
and (4) spectral clustering with normalized cuts
(scNJW).
# run algorithms
run1 = T4cluster::kmeans(data, k=4)
run2 = T4cluster::kmeanspp(data, k=4)
run3 = T4cluster::gmm(data, k=4)
run4 = T4cluster::scNJW(data, k=4, sigma = 0.1)
# visualize
par(mfrow=c(2,2))
plot(data, pch=19, xlab="", ylab="", col=run1$cluster, main="k-means")
plot(data, pch=19, xlab="", ylab="", col=run2$cluster, main="k-means++")
plot(data, pch=19, xlab="", ylab="", col=run3$cluster, main="gmm")
plot(data, pch=19, xlab="", ylab="", col=run4$cluster, main="scNJW")