lmds:
Landmark Multi-Dimensional ScalingA fast dimensionality reduction method scaleable to large numbers of samples. Landmark Multi-Dimensional Scaling (LMDS) is an extension of classical Torgerson MDS, but rather than calculating a complete distance matrix between all pairs of samples, only the distances between a set of landmarks and the samples are calculated.
library(lmds)
x <- as.matrix(iris[,1:4])
dimred <- lmds(x, ndim = 2)
qplot(dimred[,1], dimred[,2]) + labs(title = "lmds()") + theme_classic()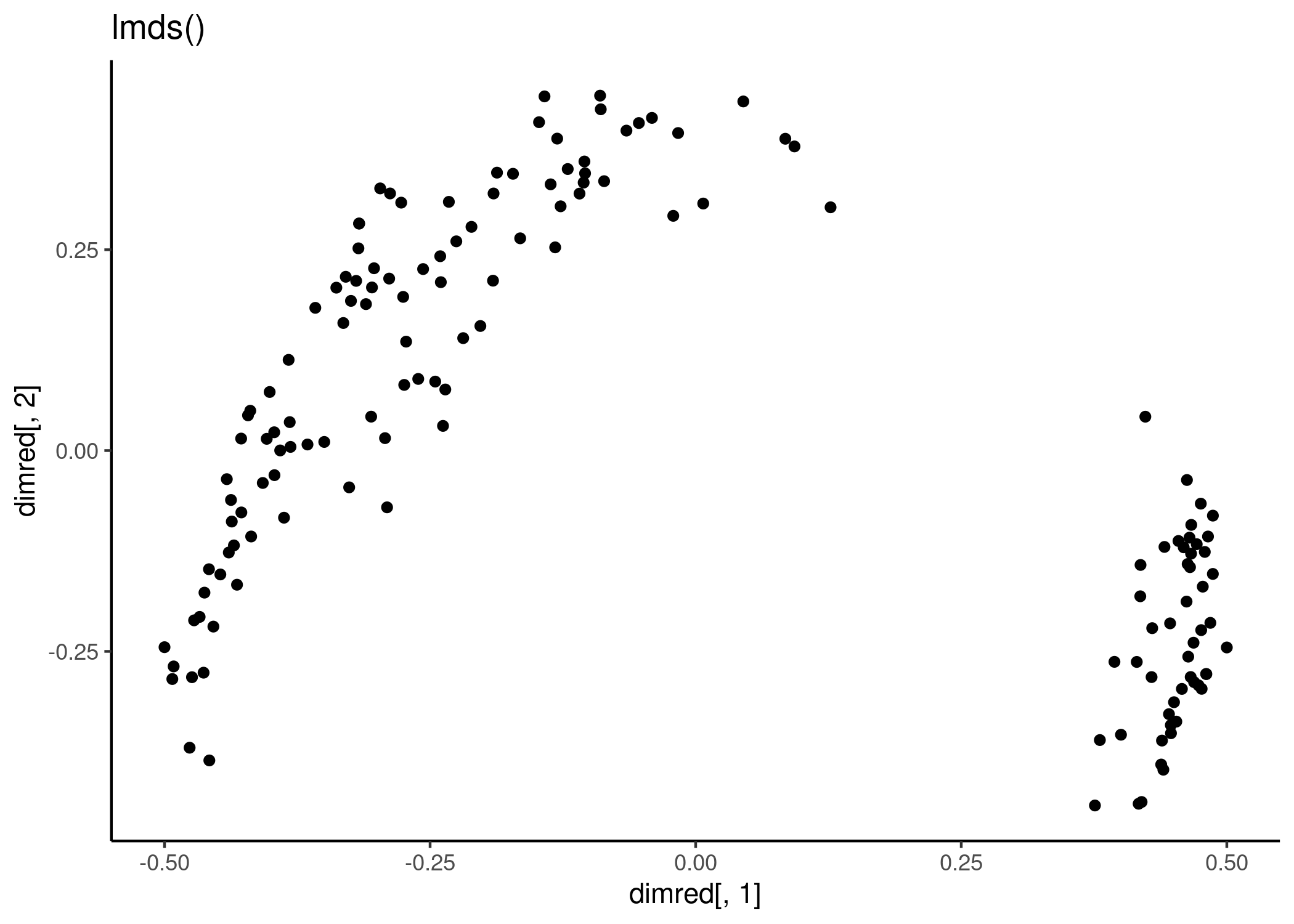
dimred <- cmdscale(dist(x))
qplot(dimred[,1], dimred[,2]) + labs(title = "cmdscale()") + theme_classic()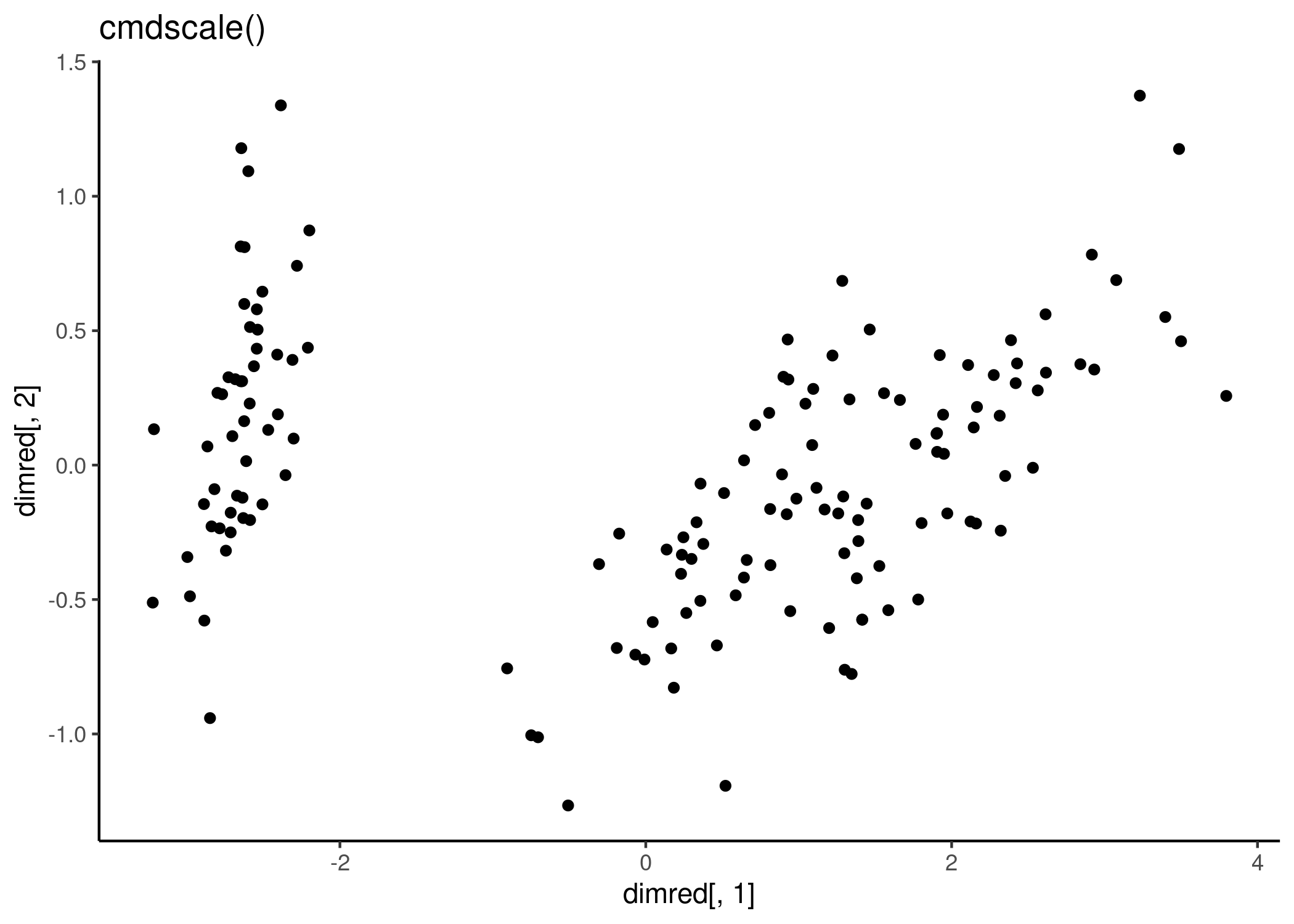
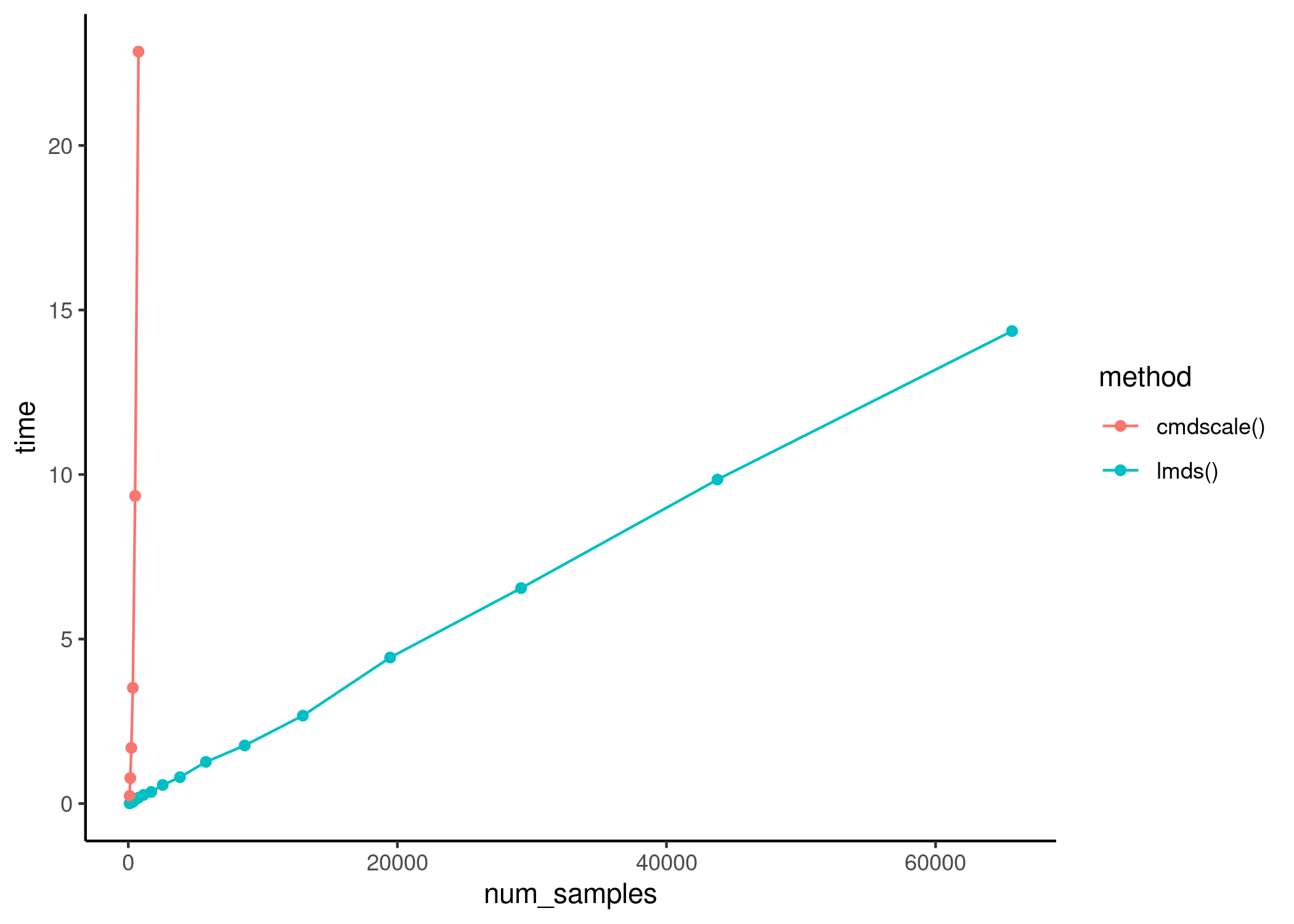
The execution time of lmds() scales linearly with
respect to the dataset size.
Check out news(package = "lmds") or NEWS.md for a full list of changes.
Initial release of lmds.